Researchers keep discovering new functions of small RNAs. For instance, they can be used as a defense mechanism against viruses or self-replicating genome invaders. These tiny pieces of RNA are often produced by a cleavage of long precursors by so called Dicer proteins. To their surprise, researchers from the University of Bern have found that some Dicers acquired a unique and as yet unknown feature that allow them to cleave the RNA precursors in a very specific way, resulting in small RNAs that work much more efficiently.
Dicer is not rolling dice
In the human cell as in society: It is all about information. Genes are read, information molecules (called messenger RNAs or short mRNA) are produced and proteins are synthesized according to the instructions. And there are a whole lot more of these messenger substances, the cellular apparatus is a huge information hub, a meticulously regulated business. And as if all these information exchanges were not complicated enough: in the past years RNA-related research has uncovered more and more evidence that regulation works on a higher level too. The information management itself is highly dynamic – information substances are being held back, they are changed, at times even shredded right away.
Wisely chosen cuts
In eukaryotes especially, the shredded information molecules, called small RNAs (sRNAs), have critical roles in development, gene expression, and genome stability. They are produced by a range of different proteins. One of the best known is called Dicer and cuts them from long double-stranded RNA templates. Up to now, Dicer was thought to be a bit of a lumberjack when it comes to tearing down RNA – just hacking it into short fragments of roughly equal size, paying no special attention to the form and content of the fragments. The proteins have never been shown to have sequence cleavage preferences, but recently a research group from the Institute of Cell Biology of the University of Bern headed by Mariusz Nowacki has found Dicer-like enzymes with truly surprising qualities. The research could lead to new insights into the regulation of cellular communication and might open up as yet uncharted opportunities for future protein engineering.
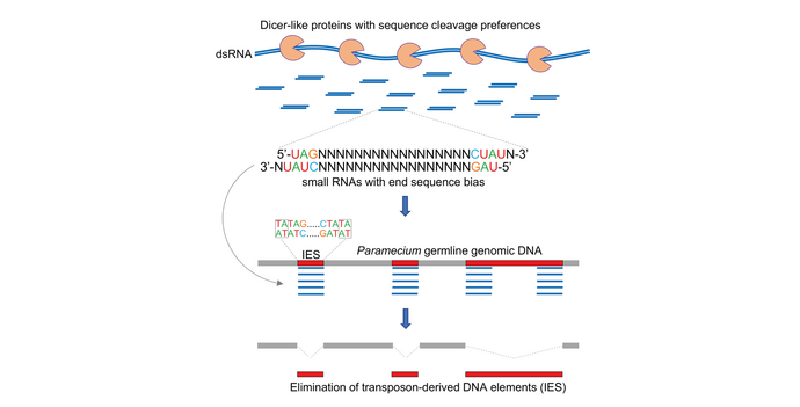
The research group, which is part of the NCCR "RNA and disease", has found that in the microorganism Paramecium, which uses small RNAs to guide the elimination of invading transposable DNA elements, some Dicer-like proteins are cutting up RNA with much greater diligence than expected. As reported in the renowned magazine Cell, Cristina Hoehener, Iris Hug, and Mariusz Nowacki could show that their Dicer-like enzymes have strict size and pronounced sequence preferences. These preferences result in the production of sRNAs precisely matching the ends of their target DNA elements, which leads the researchers to propose an as yet unknown biological role for these newly characterized enzymes and their cleavage products.
Small RNAs repair the DNA
They think that these Dicer-like proteins facilitate the elimination of unwanted DNA elements by accumulating precisely at their ends. These accumulations then trigger the elimination of the "highlighted" parts of the DNA from the genetic material. Transposons are alien parts of the genome, infiltrated into the DNA, with the ability to jump from one part of the DNA to another. This makes them especially dangerous as they can trigger all kinds of unwanted genetic effects. So an organism has every reason to get rid of these DNA segments quickly and efficiently.
Especially intriguing for the researchers is the level of precision with which these Dicer enzymes do their job. In contrast to restriction enzymes that cut only when they recognize very specific sequences, the Paramecium Dicer proteins function somewhat sloppily - Nowacki calls it a "relaxed preference". The researchers believe that a too strict preference would not be appropriate here: for the DNA repair to be effective it is important that a lot of sRNA fragments can assemble at the right segment. If Dicer would be too picky, it would not be able to produce suitable snippets for all the dangerous foreign DNA. "Fuzziness is not necessarily a bad thing", says Nowacki. Over the course of the evolution many biological processes have evened out between precision and chance - one of the reasons why cells can react with surprisingly tailor-made solutions to all kinds of different threats.
Publication details:
Christina Hoehener, Iris Hug, Mariusz Nowacki (2018): Dicer-like Enzymes with Sequence Cleavage Preferences, Cell, 173(1), 234-247 e7
https://www.sciencedirect.com/science/article/pii/S0092867418301739?via%3Dihub